5 Complete Block Designs
5.1 Introduction
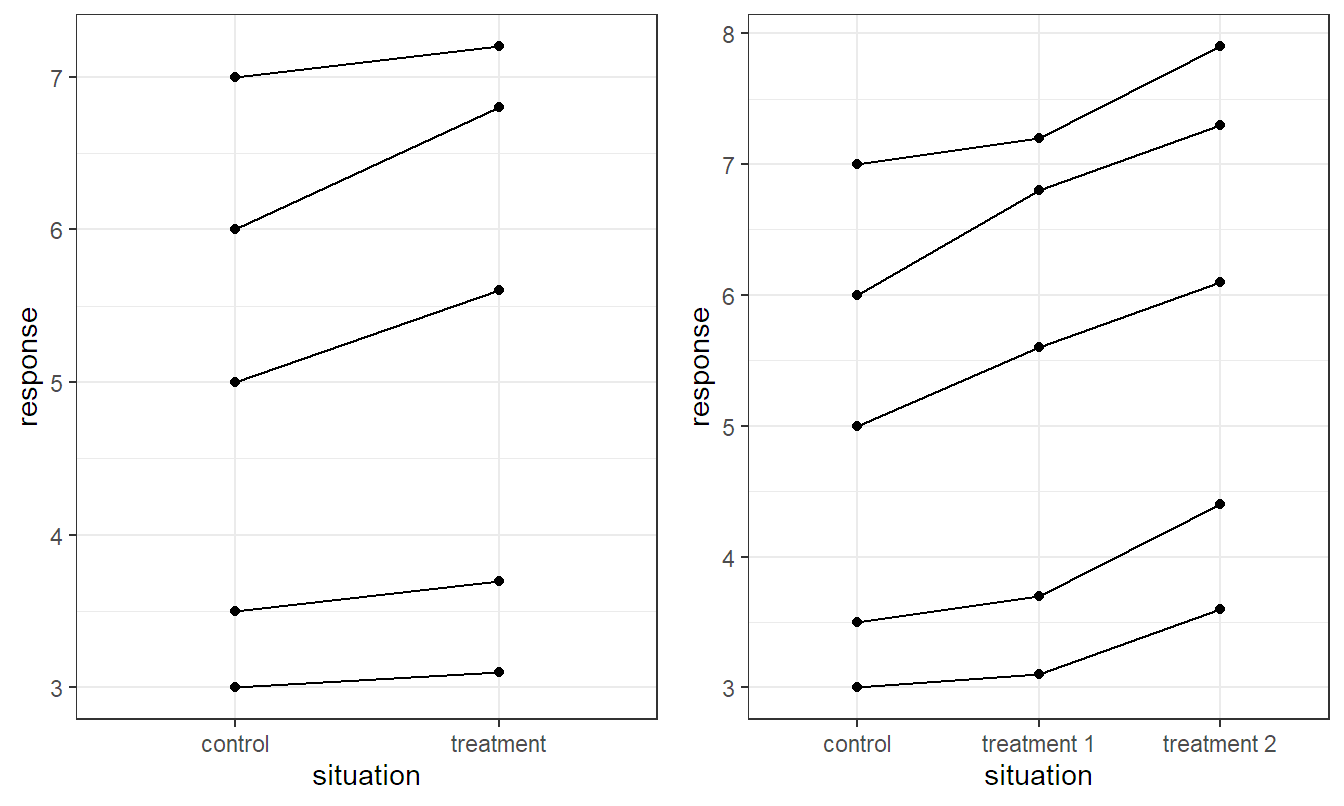
In many situations we know that our experimental units are not homogeneous. Making explicit use of the special structure of the experimental units typically helps reduce variance (“getting a more precise picture”). In your introductory course, you have learned how to apply the paired \(t\)-test. It was used for situations where two treatments were applied on the same “object” or “subject.” Think for example of applying two treatments, in parallel, on human beings (like the application of two different eye drop types, each applied in one of the two eyes). We know that individuals can be very different. Due to the fact that we apply both treatments on the same subject, we get a “clear picture” of the treatment effect within every subject by taking the difference of the response values corresponding to the two treatments. This makes the subject-to-subject variability completely disappear. We also say that we block on subjects or that an individual subject is a block. This is also illustrated in Figure 5.1 (left): The values of the same subject are connected with a line. With the help of these lines, it is obvious that the response value corresponding to treatment is larger than the value corresponding to the control group, within a subject. This would be much less obvious if the data would come from two independent groups. In this scenario, we would have to delete the lines and would be confronted with the whole variability between the different subjects.
We will now extend this way of thinking to the situation \(g > 2\), where \(g\) is the number of levels of our treatment factor (as in Chapter 2). An illustration of the basic idea can be found in Figure 5.1 (right). We simply consider situations where we have more than two levels on the \(x\)-axis. The basic idea stays the same: Values coming from the same block (here, subject) can be connected with a line. This helps both our eyes and the statistical procedure in getting a much clearer picture of the treatment effect.
5.2 Randomized Complete Block Designs
Assume that we can divide our experimental units into \(r\) groups, also known as blocks, containing \(g\) experimental units each. Think for example of an agricultural experiment at \(r\) different locations having \(g\) different plots of land each. Hence, a block is given by a location and an experimental unit by a plot of land. In the introductory example, a block was given by an individual subject.
The randomized complete block design (RCBD) uses a restricted randomization scheme: Within every block, e.g., at each location, the \(g\) treatments are randomized to the \(g\) experimental units, e.g., plots of land. In that context, location is also called the block factor. The design is called complete because we observe the complete set of treatments within every block (we will later also learn about incomplete block designs where this is not the case anymore, see Chapter 8).
Note that blocking is a special way to design an experiment, or a special “flavor” of randomization. It is not something that you use only when analyzing the data. Blocking can also be understood as replicating an experiment on multiple sets, e.g., different locations, of homogeneous experimental units, e.g., plots of land at an individual location. The experimental units should be as similar as possible within the same block, but can be very different between different blocks. This design allows us to fully remove the between-block variability, e.g., variability between different locations, from the response because it can be explained by the block factor. Hence, we get a much clearer picture for the treatment factor. The randomization step within each block makes sure that we are protected from unknown confounding variables. A completely randomized design (ignoring the blocking structure) would typically be much less efficient as the data would be noisier, meaning that the error variance would be larger. In that sense, blocking is a so-called variance reduction technique.
Typical block factors are location (see example above), day (if an experiment is run on multiple days), machine operator (if different operators are needed for the experiment), subjects, etc.
Blocking is very powerful and the general rule is, according to George Box (Box et al. 1978):
“Block what you can; randomize what you cannot.”
In the most basic form, we assume that we do not have replicates within a block. This means that we only observe every treatment once in each block.
The analysis of a randomized complete block design is straightforward. We treat the block factor as “just another” factor in our model. As we have no replicates within blocks, we can only fit a main effects model of the form \[ Y_{ij} = \mu + \alpha_i + \beta_j + \epsilon_{ij}, \] where the \(\alpha_i\)’s are the treatment effects and the \(\beta_j\)’s are the block effects with the usual side constraints. In addition, we have the usual assumptions on the error term \(\epsilon_{ij}\). According to this model, we implicitly assume that blocks only cause additive shifts. Or in other words, the treatment effects are always the same, no matter what block we consider. This assumption is usually made based on domain knowledge. If there would be an interaction effect between the block and the treatment factor, the result would be very difficult to interpret. Most often, this is due to unmeasured variables, e.g., different soil properties at different locations.
We consider an example which is adapted from Venables and Ripley (2002), the original source is Yates (1935) (we will see the full data set in Section 7.3). At six different locations (factor block), three plots of land were available. Three varieties of oat (factor variety with levels Golden.rain, Marvellous and Victory) were randomized to them, individually per location. The response was yield (in 0.25lbs per plot). A conceptual layout of the design can be found in Table 5.1.
| 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|
| Marvellous | Victory | Golden.rain | Marvellous | Marvellous | Golden.rain |
| Victory | Golden.rain | Victory | Victory | Golden.rain | Marvellous |
| Golden.rain | Marvellous | Marvellous | Golden.rain | Victory | Victory |
The data can be read as follows:
block <- factor(rep(1:6, times = 3))
variety <- rep(c("Golden.rain", "Marvellous", "Victory"), each = 6)
yield <- c(133.25, 113.25, 86.75, 108, 95.5, 90.25,
129.75, 121.25, 118.5, 95, 85.25, 109,
143, 87.25, 82.5, 91.5, 92, 89.5)
oat.variety <- data.frame(block, variety, yield)
xtabs(~ block + variety, data = oat.variety)## variety
## block Golden.rain Marvellous Victory
## 1 1 1 1
## 2 1 1 1
## 3 1 1 1
## 4 1 1 1
## 5 1 1 1
## 6 1 1 1We use the usual aov function with a model including the two main effects block and variety. It is good practice to write the block factor first; in case of unbalanced data, we would get the effect of variety adjusted for block in the sequential type I output of summary, see Section 4.2.5 and also [C]hapter Chapter 8].
## Df Sum Sq Mean Sq F value Pr(>F)
## block 5 3969 794 5.28 0.012
## variety 2 447 223 1.49 0.272
## Residuals 10 1503 150We first focus on the p-value of the treatment factor variety. Although we used a randomized complete block design, we cannot reject the null hypothesis that there is no overall effect of variety (a reason might be low power, as we only have 10 degrees of freedom left for the error term). Typically, we are not inspecting the p-value of the block factor block. There is some historic debate why we should not do this, mainly because of the fact that we did not randomize blocks to experimental units. In addition, we already knew (or hoped) beforehand that blocks are different. Hence, such a finding would not be of great scientific relevance. However, we can do a quick check to verify whether blocking was efficient or not. We would like the block factor to explain a lot of variation, hence if the mean square of the block factor is larger than the error mean square \(MS_E\) we conclude that blocking was efficient (compared to a completely randomized design). Here, this is the case as \(794 > 150\). See Kuehl (2000) for more details and a formal definition of the relative efficiency which compares the efficiency of a randomized complete block design to a completely randomized design. If blocking was not efficient, we would still leave the block factor in the model (as the model must follow the design that we used), but we might plan not to use blocking in a future similar experiment because it didn’t help reduce variance and only cost us degrees of freedom.
Instead of a single treatment factor, we can also have a factorial treatment structure within every block. Think for example of a design as outlined in Table 5.2.
| \(1\) | \(2\) | \(3\) | \(\cdots\) |
|---|---|---|---|
| \(A_1B_2\) | \(A_2B_2\) | \(A_2B_1\) | \(\cdots\) |
| \(A_2B_1\) | \(A_2B_1\) | \(A_1B_2\) | \(\cdots\) |
| \(A_2B_2\) | \(A_1B_2\) | \(A_1B_1\) | \(\cdots\) |
| \(A_1B_1\) | \(A_1B_1\) | \(A_2B_2\) | \(\cdots\) |
In R, we would model this as y ~ Block + A * B. In such a situation, we can actually test the interaction between \(A\) and \(B\) even if every level combination of \(A\) and \(B\) appears only once in every block. Why? Because we have multiple blocks, we have multiple observations for every combination of the levels of \(A\) and \(B\). Of course, the three-way interaction cannot be added to the model.
Interpretation of the coefficients of the corresponding models, residual analysis, etc. is done “as usual.” The only difference is that we do not test the block factor for statistical significance, but for efficiency.
5.3 Nonparametric Alternatives
For a complete block design with only one treatment factor and no replicates, there is a rank sum based test, the so-called Friedman rank sum test which is implemented in function friedman.test. Among others, it also has a formula interface, where the block factor comes after the symbol “|”, i.e., for the oat example the formula would be yield ~ variety | block:
friedman.test(yield ~ variety | block, data = oat.variety)##
## Friedman rank sum test
##
## data: yield and variety and block
## Friedman chi-squared = 2.3, df = 2, p-value = 0.35.4 Outlook: Multiple Block Factors
We can also block on more than one factor. A special case is the so-called Latin Square design where we have two block factors and one treatment factor having \(g\) levels each (yes, all of them!). Hence, this is a very restrictive assumption. Consider the layout in Table 5.3 where we have a block factor with levels \(R_1\) to \(R_4\) (“rows”), another block factor with levels \(C_1\) to \(C_4\) (“columns”) and a treatment factor with levels \(A\) to \(D\) (a new notation as now the letter is actually the level of the treatment factor). In a Latin Square design, each treatment (Latin letters) appears exactly once in each row and once in each column. A Latin Square design blocks on both rows and columns simultaneously. We also say it is a row-column design. If we ignore the columns of a Latin Square designs, the rows form an RCBD; if we ignore the rows, the columns form an RCBD.
| \(C_1\) | \(C_2\) | \(C_3\) | \(C_4\) | |
|---|---|---|---|---|
| \(R_1\) | \(A\) | \(B\) | \(C\) | \(D\) |
| \(R_2\) | \(B\) | \(C\) | \(D\) | \(A\) |
| \(R_3\) | \(C\) | \(D\) | \(A\) | \(B\) |
| \(R_4\) | \(D\) | \(A\) | \(B\) | \(C\) |
For example, if we have a factory with four machine types (treatment factor with levels \(A\) to \(D\)) such that each of four operators can only operate one machine on a single day, we could perform an experiment on four days and block on days (\(C_1\) to \(C_4\)) and operators (\(R_1\) to \(R_4\)) using a Latin Square design as shown in Table 5.3.
By design, a Latin Square with a treatment factor with \(g\) levels uses (only) \(g^2\) experimental units. Hence, for small \(g\), the degrees of freedom of the error term can be very small, see Table 5.4, leading to typically low power.
| \(g\) | Error Degrees of Freedom |
|---|---|
| 3 | 2 |
| 4 | 6 |
| 5 | 12 |
| 6 | 20 |
We can create a (random) Latin Square design in R for example with the function design.lsd of the package agricolae (de Mendiburu 2020).
library(agricolae)
design.lsd(LETTERS[1:4])$sketch## [,1] [,2] [,3] [,4]
## [1,] "A" "C" "B" "D"
## [2,] "C" "A" "D" "B"
## [3,] "D" "B" "A" "C"
## [4,] "B" "D" "C" "A"To analyze data from such a design, we use the main effects model \[ Y_{ijk} = \mu + \alpha_i + \beta_j + \gamma_k + \epsilon_{ijk}. \] Here, the \(\alpha_i\)’s are the treatment effects and \(\beta_j\) and \(\gamma_k\) are the row- and column-specific block effects with the usual side constraints.
The design is balanced having the effect that our usual estimators and sums of squares are “working.” In R, we would use the model formula y ~ Block1 + Block2 + Treat. We cannot fit a more complex model, including interaction effects, here because we do not have the corresponding replicates.
Multiple Latin Squares can also be combined or replicated. This allows for more flexibility. In that sense, Latin Square designs are useful building blocks of more complex designs, see for example Kuehl (2000).
With patients, it is common that one is not able to apply multiple treatments in parallel, but in a sequential manner over multiple time periods (think of comparing different painkillers: a different painkiller per day). In such a situation, we would like to block on patients (“as usual”) but also on time periods, because there might be something like a learning or a fatigue effect over time. Such designs are called crossover designs, a standard reference is Jones and Kenward (2014). If the different periods are too close to each other, so-called carryover effects might be present: the medication of the previous period might still have an effect on the current period. Such effects can be incorporated into the model (if this is part of the research question). Otherwise, long enough washout periods should be used.